Gene Regulatory Networks
We build computational tools to map and model the networks of genes that control cell identity.
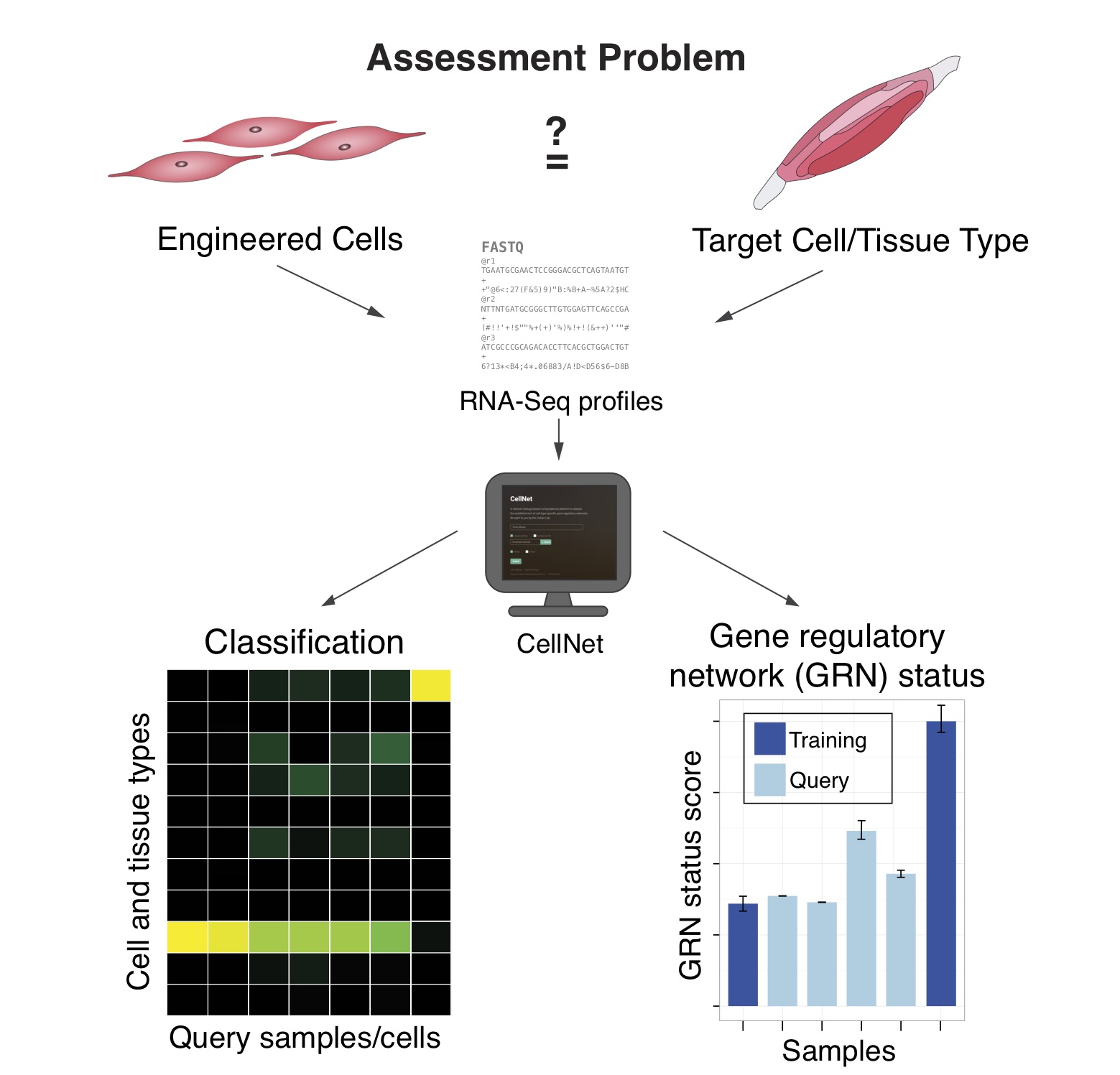
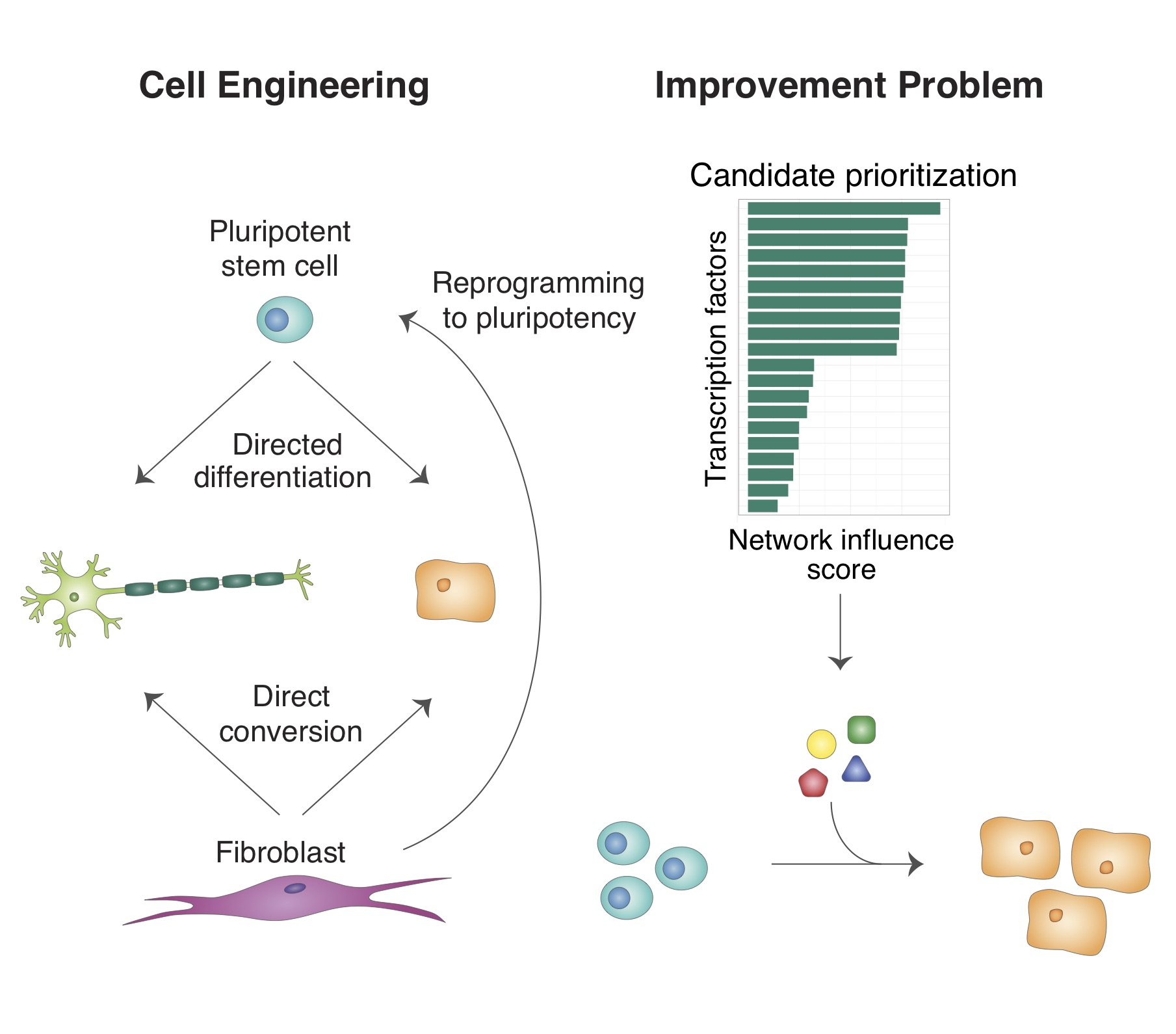
Gene regulatory networks (GRNs) define the complete set of regulatory relationships among genes and gene products in a cell. These networks govern a cell’s transcriptional output and act as the molecular blueprint of cell-type identity. We develop algorithms to reconstruct GRNs from genomic data, measure how they are established during development, infer their dynamics, and model regulatory interactions between cells — with a particular focus on synovial joint development.
 Johns Hopkins University
Johns Hopkins University